The Duke Campus Farm typically sees more visitors than usual on Fridays, when it holds Community Work Days and welcomes students, faculty, and community members to help run tasks and learn more about its sustainable agriculture practices.
However, this particular Friday, April 11, was a bit special. Instead of us volunteers driving wheelbarrows back and forth, mulching or weeding, several members of the Vilgalys Lab at Duke instructed participants on how to grow our own mushrooms.
Shiitake Logs
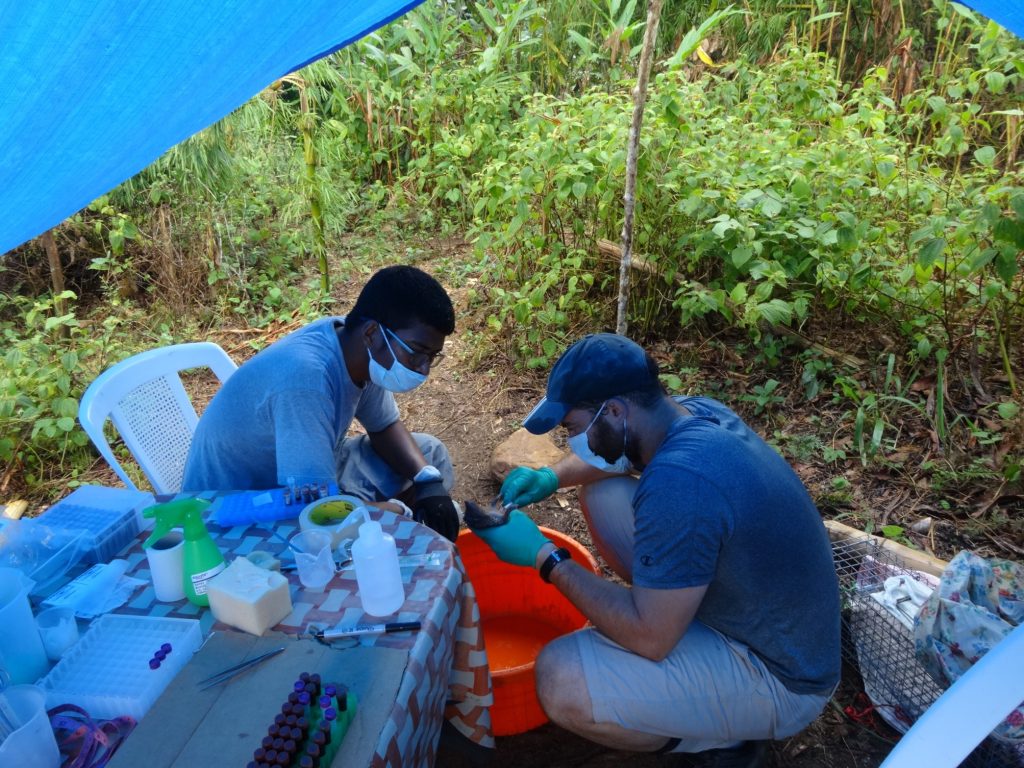
The process begins with inoculation: placing the mushroom spawn into substrate, which provides it a suitable environment to live in. It turns out there’s two common ways to do this, so we split ourselves into two groups. I headed over to the right side first, where Dr. Rytas Vilgalys awaited us with a bag of shittake spawn. A Duke biology professor, Vilgalys is an expert on mycology, the study of fungi. He was joined by undergraduate senior Mira Polishook, who is both part of the Vilgalys lab and the farm’s programming crew.
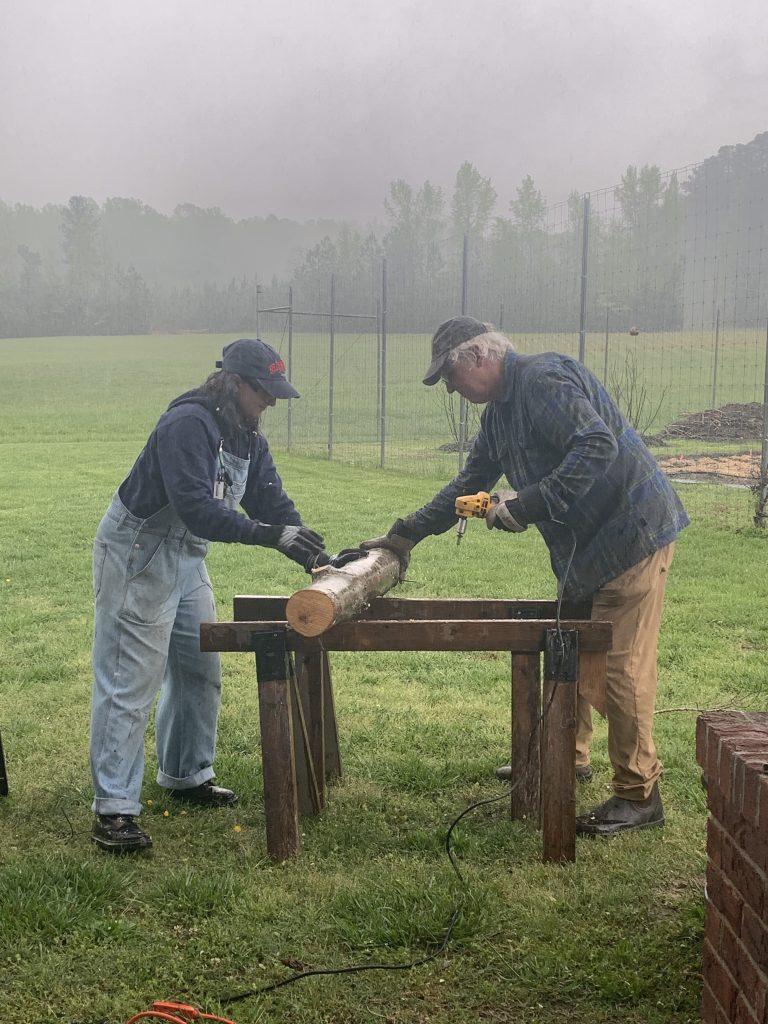

They introduced us to our substrate, sweetgum logs, which sat neatly stacked in a pile to the side. Like these, suitable logs will be from hardwood trees and recently cut. (Many softwoods, like pines, have anti-fungal properties, and old logs are often already colonized by other fungi.)
The first task was to create holes in the logs for the spawn. After a quick demonstration, Vilgalys handed off the drill, encouraging those of us who hadn’t used power tools before to do so.
A friend and I quickly formed a system–I drilled holes as uniformly as I could while they rotated the log after every row. Though the day was dreary and wet around us, it soon became lively with movement and chatter underneath the pavilion as everyone began carrying out their roles.
After a few turns, I relinquished the drill to someone else and instead picked up the inoculation thumb tool. With narrow metal cylinders, these made it easy to pick up an appropriate amount of our sawdust spawn and release it into the holes we drilled. (These aren’t always necessary with other methods; plug spawn comes in pellet form and can be simply placed into the holes by hand.)
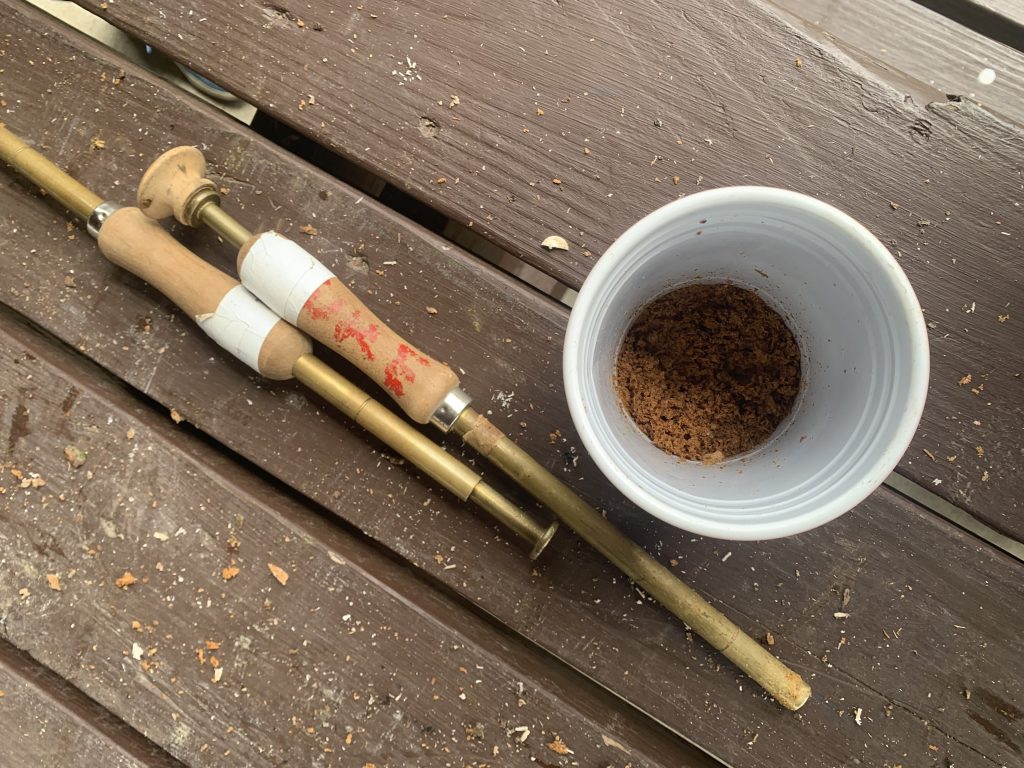




A few tables over, others did the finishing touches via paintbrush. Fungi require a moist environment to grow, meaning mycelium in open holes are at risk of drying out. Dripping hot wax over the holes seals moisture inside and prevent insects from getting inside, until the wax eventually breaks off as the fungi grows outward.
Oyster Bags

On the other side of the pavilion, Duke mycologist Khalid Hameed was leading workshop participants through creating oyster mushroom bags, using a different type of substrate and spawn.
“Substrate is any lignin or cellulose. This is wheat straw. You can use rice straw, you can use peanut straw…any lignin, cellulose material,” Hameed said. From the bags that he had prepared for us, we took fistfuls of damp wheat straw and created the first layer in our own clear bags. Over this went a sprinkle of grain spawn. Repeatedly, we built up layers of straw and spawn until our bags were full.
“The first stage of this, we call pinhair. It [the fungus] makes little tiny pinhairs,” Hameed said. For a couple of weeks, there’s essentially no maintenance required. As long as there’s some humidity in the air, the bag can sit in a dark cellar or room. “If the moisture is not enough, those pinhairs then will dry.”

This method is relatively quick, and we soon arrived at Hameed’s final instruction. Using a small razor edge, we pricked the bags evenly all over, creating small tears in the shape of crosses. The edible parts of fungi–the outward visible parts–are called “fruit,” and it’s out of these holes that the oysters will eventually fruit.
Good Mushrooms Come to Those That Wait
By the end of the workshop, the evidence of our work surrounded us: the tables lay covered in sawdust, wax drops, and stray wheatgrass.
Now, we wait for the (literal) fruit of our labor. I’ll be checking on my oyster bag, which will appear more and more white as the fungus colonizes the straw. These will only need around three weeks to begin fruiting, at which point they need to be moved into a room with light. However, the logs we inoculated likely won’t fruit for at least a year. When that time approaches, the fungi will need disturbing or “shocking”, which can involve soaking the logs in water, knocking them down, or as they do in Japan, hitting them with hammers. It’s theorized that this shocking promotes rapid growth by stimulating natural conditions like the falling of a tree or change of weather.
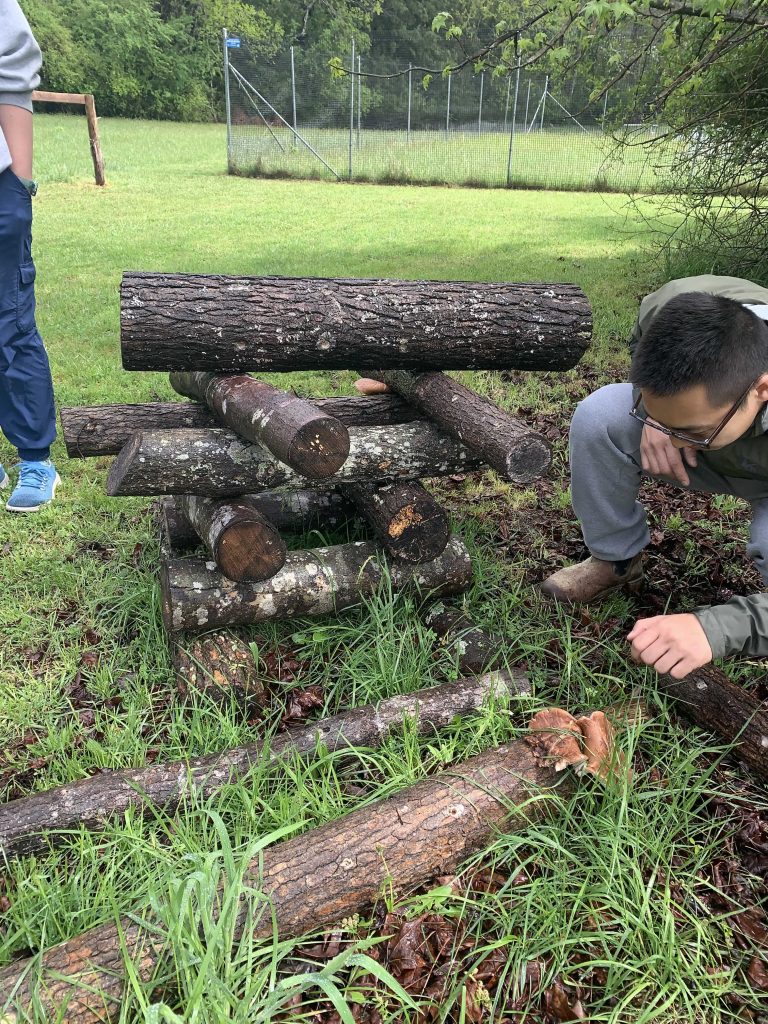



Otherwise, nothing more that needs to be done. Once inoculated, logs can continue to fruit year after year. The logs above, which were inoculated last year at Duke Campus Farm, have since fruited, though not to their fullest extent. For the work of one afternoon, the payoff is significant–an easy, sustainable way to farm food.

By Crystal Han, Class of 2028